I did the png conversion but skipped the regularization weight and data augmentation adjustments, and now it actually works with Mapillary Vistas. For completeness, I will list all necessary steps in this message, although most of them are copied from my previous messages.
- Image and label resizing using Python:
#!/usr/bin/env python3
import os
import cv2 as cv
import numpy as np
from PIL import Image
TRAIN_IMAGE_DIR = '/path/to/mapillary_vistas_2/training/images'
TRAIN_LABEL_DIR = '/path/to/mapillary_vistas_2/training/v2.0/labels'
VAL_IMAGE_DIR = '/path/to/mapillary_vistas_2/validation/images'
VAL_LABEL_DIR = '/path/to/mapillary_vistas_2/validation/v2.0/labels'
image_dirs = [TRAIN_IMAGE_DIR, VAL_IMAGE_DIR]
label_dirs = [TRAIN_LABEL_DIR, VAL_LABEL_DIR]
target_size = (512, 512)
dir_suffix = '_{}x{}'.format(*target_size)
def resize_image(filename_old, filename_new):
image = cv.imread(filename_old, cv.IMREAD_UNCHANGED)
image = cv.resize(image, dsize=target_size, interpolation=cv.INTER_AREA)
cv.imwrite(filename_new, image)
def resize_label(filename_old, filename_new):
image = np.array(Image.open(filename_old).convert('P'))
image = cv.resize(image, dsize=target_size, interpolation=cv.INTER_NEAREST)
cv.imwrite(filename_new, image)
def process_dir(dirname_old, resize_fun):
dirname_new = os.path.normpath(dirname_old) + dir_suffix
if os.path.exists(dirname_new):
print('{} already exists, skipping'.format(dirname_new))
return
os.mkdir(dirname_new)
files = os.listdir(dirname_old)
for f in files:
filename_old = os.path.join(dirname_old, f)
filename_new = os.path.join(dirname_new, f)
resize_fun(filename_old, filename_new)
for d in image_dirs:
process_dir(d, resize_image)
for d in label_dirs:
process_dir(d, resize_label)
- Jpg-to-png conversion (this could probably be easily done by modifying the Python script above, but since I had already run it, I used
convertinstead):
mkdir /path/to/mapillary_vistas_2/training/images_512x512_png
for f in /path/to/mapillary_vistas_2/training/images_512x512/*.jpg
do
convert "$f" /path/to/mapillary_vistas_2/training/images_512x512_png/$(basename "$f" .jpg).png
done
mkdir /path/to/mapillary_vistas_2/validation/images_512x512_png
for f in /path/to/mapillary_vistas_2/validation/images_512x512/*.jpg
do
convert "$f" /path/to/mapillary_vistas_2/validation/images_512x512_png/$(basename "$f" .jpg).png
done
- Docker launch:
docker run -it --gpus all \
-v /path/to/mapillary_vistas_2:/mapillary_vistas_2:ro \
-v /path/to/my_folder:/workspace/my_folder \
nvcr.io/nvidia/tlt-streamanalytics:v3.0-py3
- Model download (note that this is using TAO although the docker image is TLT):
ngc registry model download-version nvidia/tao/pretrained_semantic_segmentation:resnet18
- Spec file and training command:
model.txt (19.5 KB)
unet train \
-e /workspace/my_folder/model.txt \
-m /workspace/pretrained_semantic_segmentation_vresnet18/resnet_18.hdf5 \
-r /workspace/my_folder/output \
-k my_key
This is the training loss:
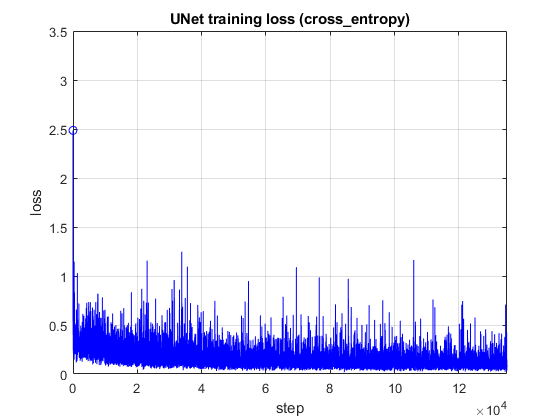
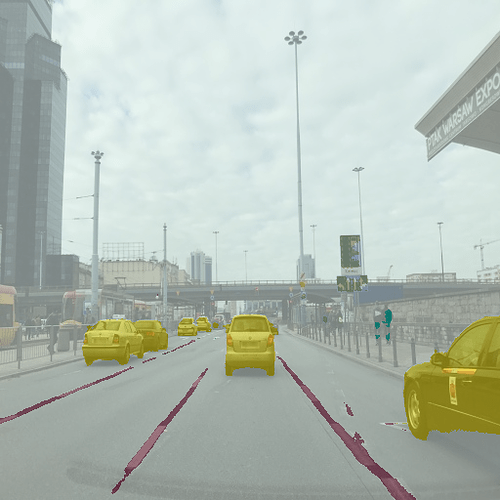
And this is the inference result:
Things to note:
- Automatic mixed precision was not used
- All images were png
- All image aspect ratios matched the network input aspect ratio
- All images actually had the same size as the network input, but I assume this is not necessary
Also, when resizing labels, it is important to make sure that the output is in single-channel format, and that the interpolation method is chosen such that no spurious labels are introduced. The above Python scripts does the resizing properly.
I won’t mark this as solved yet, because I want to train the network using the BDD100K dataset which was my original goal. I also want to run inference with different image sizes. Next, I will investigate whether I can achieve these goals.
